Application talk
Abstract
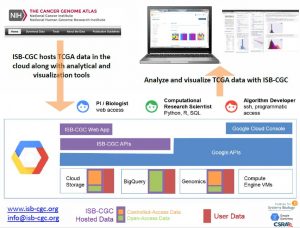 The ISB Cancer Genomics Cloud (ISB-CGC) is one of three pilot projects funded by the National Cancer Institute with the goal of democratizing access to The Cancer Genome Atlas (TCGA) data by substantially lowering the barriers to accessing and computing over this rich dataset. The ISB-CGC is a cloud-based platform that serves as a large-scale data repository for TCGA data, while also providing the computational infrastructure and interactive exploratory tools necessary to carry out cancer genomics research at unprecedented scales. The ISB-CGC facilitates collaborative research by allowing scientists to share data, analyses, and insights in a cloud environment. The ISB-CGC team includes scientists and engineers from the Institute for Systems Biology (ISB), Google, and CSRA. Goals The ISB-CGC aims to serve the needs of a broad range of cancer researchers: from scientists or clinicians who prefer to use an interactive web-based application to access and explore the rich TCGA dataset; to computational scientists who want to write their own custom scripts using languages such as R or Python, accessing the data through APIs; to algorithm developers who want to spin up thousands of virtual machines to analyze hundreds of terabytes of sequence data. The ISB-CGC allows scientists to interactively define and compare cohorts, examine the underlying molecular data for specific genes or pathways of interest, and share insights with collaborators around the globe. As part of our on-going community evaluation phase, ISB-CGC users may apply for Google Cloud Platform credits that can be used to upload their own datasets into Google Cloud Storage, and to perform analyses using existing or custom pipelines. Technology and data The ISB-CGC provides interactive and programmatic access to the TCGA data, leveraging many Google Cloud Platform technologies including BigQuery, Compute Engine, and Google Genomics. Open-access clinical and biospecimen information for all TCGA patients and samples, combined with the normalized and summarized ‘Level-3’ TCGA data and a variety of genomic reference and platform-annotation sources are stored in BigQuery, enabling fast SQL-like queries against the entire dataset. Controlled-access DNA and RNA sequence data are available to dbGaP-authorized users in the original BAM and FASTQ file formats, and using the Global Alliance for Genomics and Health (GA4GH) API. To supplement the TCGA BigQuery tables and expand the range of analyses that can be performed using BigQuery, the ISB-CGC team is also making reference tables based on other public data available such as the GENCODE gene set, the miRTarBase database of microRNA-target interactions, and the Illumina DNA Methylation platform annotation. Many other publicly-available BigQuery tables exist already, including variants from the 1,000 Genomes and Illumina Platinum Genomes projects, the Tute Genomics Annotation table, and the hg19 reference genome. In the summer of 2016, the ISB-CGC platform will also provide access to the TCGA low-level sequence data in a more “cloud-aware” form, i.e. in Google Genomics. Google Genomics includes an implementation of the GA4GH API, layered on top of Spanner and Bigtable (highly-scalable, massively parallel noSQL database technologies). Many common tasks involving DNA or RNA sequence data can be divided into millions of independent steps, and with the TCGA data available in this form, computational users will be able to implement massively parallel methods using frameworks such as Google Cloud Dataflow, Apache Hadoop, or Apache Spark. Over the lifetime of the ISB-CGC platform, we will continue to roll out new features in our web-based user-interface, while continuing to add computational examples to our public github. Upcoming interactive features will include integration with the Integrative Genomics Viewer (IGV) and the Next Generation Clustered Heat Maps (NG-CHM). Additionally, the ISB-CGC team is creating BigQuery tables with data from large cancer cell line screening efforts, such as the Cancer Cell Line Encyclopedia (CCLE) and Genomics of Drug Sensitivity in Cancer (GDSC). Reproducibility and scalability The existence of TCGA and other data in appropriate cloud data formats enables the development and deployment of integrative workflows for rapid, flexible and reproducible analyses. The use of virtual machines and containerized software tools will minimize difficulties that frequently arise due to differences between local hardware and software configurations. In addition, cloud-based data repositories eliminate the need for each researcher to download and store their own copy of the data. Importantly, computational analysis workflows can easily be shared with the research community. The ISB-CGC team has created a public GitHub repository containing R and Python scripts to demonstrate many example analyses using the TCGA BigQuery tables. These resources allow computational biologists to work in a familiar programming environment while providing a robust, scalable computational infrastructure that supports flexibility and reproducibility. Further information This project has been funded in whole with Federal funds from the National Cancer Institute, National Institutes of Health, Department of Health and Human Services, under Contract No. HHSN261201400007C. If you are interested in learning more about the ISB-CGC or would like to propose specific scientific use-cases to our development team, please visit us at www.isb-cgc.org.
The ISB Cancer Genomics Cloud (ISB-CGC) is one of three pilot projects funded by the National Cancer Institute with the goal of democratizing access to The Cancer Genome Atlas (TCGA) data by substantially lowering the barriers to accessing and computing over this rich dataset. The ISB-CGC is a cloud-based platform that serves as a large-scale data repository for TCGA data, while also providing the computational infrastructure and interactive exploratory tools necessary to carry out cancer genomics research at unprecedented scales. The ISB-CGC facilitates collaborative research by allowing scientists to share data, analyses, and insights in a cloud environment. The ISB-CGC team includes scientists and engineers from the Institute for Systems Biology (ISB), Google, and CSRA. Goals The ISB-CGC aims to serve the needs of a broad range of cancer researchers: from scientists or clinicians who prefer to use an interactive web-based application to access and explore the rich TCGA dataset; to computational scientists who want to write their own custom scripts using languages such as R or Python, accessing the data through APIs; to algorithm developers who want to spin up thousands of virtual machines to analyze hundreds of terabytes of sequence data. The ISB-CGC allows scientists to interactively define and compare cohorts, examine the underlying molecular data for specific genes or pathways of interest, and share insights with collaborators around the globe. As part of our on-going community evaluation phase, ISB-CGC users may apply for Google Cloud Platform credits that can be used to upload their own datasets into Google Cloud Storage, and to perform analyses using existing or custom pipelines. Technology and data The ISB-CGC provides interactive and programmatic access to the TCGA data, leveraging many Google Cloud Platform technologies including BigQuery, Compute Engine, and Google Genomics. Open-access clinical and biospecimen information for all TCGA patients and samples, combined with the normalized and summarized ‘Level-3’ TCGA data and a variety of genomic reference and platform-annotation sources are stored in BigQuery, enabling fast SQL-like queries against the entire dataset. Controlled-access DNA and RNA sequence data are available to dbGaP-authorized users in the original BAM and FASTQ file formats, and using the Global Alliance for Genomics and Health (GA4GH) API. To supplement the TCGA BigQuery tables and expand the range of analyses that can be performed using BigQuery, the ISB-CGC team is also making reference tables based on other public data available such as the GENCODE gene set, the miRTarBase database of microRNA-target interactions, and the Illumina DNA Methylation platform annotation. Many other publicly-available BigQuery tables exist already, including variants from the 1,000 Genomes and Illumina Platinum Genomes projects, the Tute Genomics Annotation table, and the hg19 reference genome. In the summer of 2016, the ISB-CGC platform will also provide access to the TCGA low-level sequence data in a more “cloud-aware” form, i.e. in Google Genomics. Google Genomics includes an implementation of the GA4GH API, layered on top of Spanner and Bigtable (highly-scalable, massively parallel noSQL database technologies). Many common tasks involving DNA or RNA sequence data can be divided into millions of independent steps, and with the TCGA data available in this form, computational users will be able to implement massively parallel methods using frameworks such as Google Cloud Dataflow, Apache Hadoop, or Apache Spark. Over the lifetime of the ISB-CGC platform, we will continue to roll out new features in our web-based user-interface, while continuing to add computational examples to our public github. Upcoming interactive features will include integration with the Integrative Genomics Viewer (IGV) and the Next Generation Clustered Heat Maps (NG-CHM). Additionally, the ISB-CGC team is creating BigQuery tables with data from large cancer cell line screening efforts, such as the Cancer Cell Line Encyclopedia (CCLE) and Genomics of Drug Sensitivity in Cancer (GDSC). Reproducibility and scalability The existence of TCGA and other data in appropriate cloud data formats enables the development and deployment of integrative workflows for rapid, flexible and reproducible analyses. The use of virtual machines and containerized software tools will minimize difficulties that frequently arise due to differences between local hardware and software configurations. In addition, cloud-based data repositories eliminate the need for each researcher to download and store their own copy of the data. Importantly, computational analysis workflows can easily be shared with the research community. The ISB-CGC team has created a public GitHub repository containing R and Python scripts to demonstrate many example analyses using the TCGA BigQuery tables. These resources allow computational biologists to work in a familiar programming environment while providing a robust, scalable computational infrastructure that supports flexibility and reproducibility. Further information This project has been funded in whole with Federal funds from the National Cancer Institute, National Institutes of Health, Department of Health and Human Services, under Contract No. HHSN261201400007C. If you are interested in learning more about the ISB-CGC or would like to propose specific scientific use-cases to our development team, please visit us at www.isb-cgc.org.
Authors
Theo Knijnenburg, Institute for Systems Biology, United States
Ilya Shmulevich, Institute for Systems Biology, United States
Sheila Reynolds, Institute for Systems Biology, United States
Phyliss Lee, Institute for Systems Biology, United States
Michael Miller, Institute for Systems Biology, United States
Kelly Iverson, Institute for Systems Biology, United States
Abigail Hahn, Institute for Systems Biology, United States
Zack Rodebaugh, Institute for Systems Biology, United States
Kalle Leinonen, Institute for Systems Biology, United States
Dave Gibbs, Institute for Systems Biology, United States
Varsha Dhankani, Institute for Systems Biology, United States
Jonathan Bingham, Google Genomics, United States
Nicole Deflaux, Google Genomics, United States
Matt Bookman, Google Genomics, United States
David Pot, CSRA, United States
